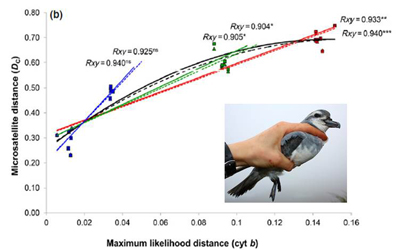
Microsatellite loci are ideal for testing hypotheses relating to genetic segregation at fine spatio-temporal scales. They are also conserved among closely related species, making them potentially useful for clarifying interspecific relationships between recently diverged taxa. However, mutations at primer binding sites may lead to increased nonamplification, or disruptions that may result in decreased polymorphism in nontarget species. Furthermore, high mutation rates and constraints on allele size may also with evolutionary time, promote an increase in convergently evolved allele size classes, biasing measures of interspecific genetic differentiation. Here, next-generation sequencing was used to develop microsatellite markers from a shotgun genome sequence of the sub-Antarctic seabird, the thin-billed prion (Pachyptila belcheri), and tested for cross-species amplification in other Pachyptila and related sub-Antarctic species. It was found that heterozygosity decreased and the proportion of nonamplifying loci increased with phylogenetic distance from the target species. Surprisingly, species trees estimated from interspecific FST provided better approximations of mtDNA relationships among the studied species than those estimated using DC, even though FST was more affected by null alleles. A significantly nonlinear second order polynomial relationship between microsatellite and mtDNA distances was observed. Authors propose that the loss of linearity with increasing mtDNA distance stems from an increasing proportion of homoplastic allele size classes that are identical in state, but not identical by descent. Therefore, despite high cross-species amplification success and high polymorphism among the closely related Pachyptila species, we caution against the use of microsatellites in phylogenetic inference among distantly related taxa. informacion[at]ebd.csic.es Moodley et al (2015) Evolutionary factors affecting the cross-species utility of newly developed microsatellite markers in seabirds Mol Ecol Res DOI: 10.1111/1755-0998.12372
http://onlinelibrary.wiley.com/doi/10.1111/1755-0998.12372/abstract